Fine-Tuning ProGen2 Protein Language Model
Apr 15, 2025
·
1 min read
Fine-tuned ProGen2 transformer model on PFAM protein families to generate biologically valid sequences. Achieved 93%+ motif retention rates through targeted hyperparameter optimization and attention mechanism analysis.
Overview
This project demonstrates transfer learning applied to protein sequence generation, fine-tuning a large pre-trained transformer model (ProGen2) on specific protein families to generate novel sequences while preserving critical biological motifs.
Technical Implementation
Model Training:
- Fine-tuned ProGen2 on three PFAM families (PF00257, PF00069, PF00072) using GPU cluster
- Built both single-family and multi-family models achieving 20% and 25% average sequence identity
- Training time: ~3 hours per epoch on distributed GPUs
Optimization:
- Reduced perplexity from 1.72 to 1.30 on PF00257 through hyperparameter tuning
- Systematic exploration of learning rates, batch sizes, and warmup schedules
Validation:
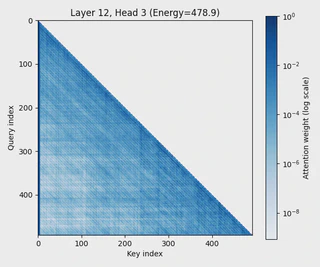
- Applied attention heatmap visualization to understand learned dependencies
- HMMer analysis confirmed 93.9% motif retention for single-family model
- Cross-family validation showed 93% retention for PF00072
Key Technologies
- PyTorch for model implementation
- Hugging Face Transformers for ProGen2 architecture
- GPU cluster computing for distributed training
- HMMer for biological sequence validation
Impact
Successfully demonstrated that careful fine-tuning of large language models can generate biologically plausible protein sequences, with validation metrics confirming preservation of critical functional motifs.